AI Antibiotic Discovery: How 46 Billion Molecules Led to a New MRSA Candidate
The idea that artificial intelligence could help discover entirely new antibiotics once sounded like a distant promise. Today, it’s starting to look much more tangible. A recent study published in Molecular Systems Biology shows just how far this field has progressed — with an AI system scanning a staggering 46 billion possible molecules and identifying a promising candidate against one of the most stubborn bacterial threats: MRSA.
But beyond the headline numbers, what really matters is how this system works — and why it represents a meaningful shift in how we search for new drugs.
This impressive achievement in using AI for antibiotic discovery highlights a larger trend, as Neural Networks in Medicine: AI Transforming Healthcare across diagnostics, drug discovery, and beyond.
Table of Contents
From Data to Molecules: What SyntheMol-RL Actually Does
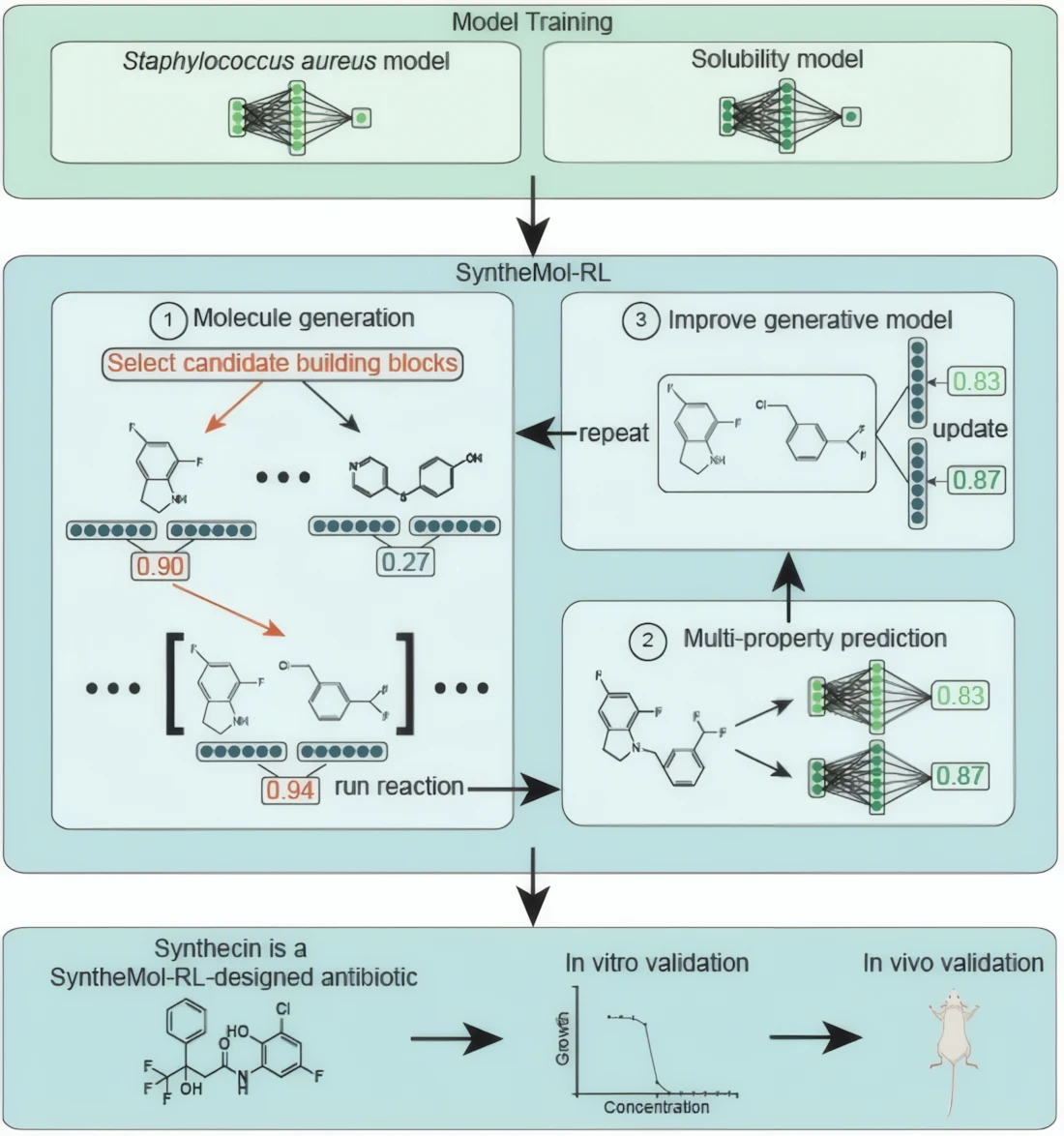
At the center of the research is SyntheMol-RL, an AI system developed by scientists from McMaster University and Stanford. It builds on an earlier framework but introduces a key upgrade: reinforcement learning. That may sound technical, but the idea is surprisingly intuitive — the system learns by optimizing outcomes based on multiple goals at once.
In this case, researchers trained it to focus on two crucial properties. First, the molecule had to be effective against Staphylococcus aureus. Second, it needed to be soluble in water — a seemingly simple requirement that often becomes a major bottleneck in drug development. After all, a compound that works in theory but fails in real biological conditions is essentially useless.
What makes SyntheMol-RL particularly interesting is that it doesn’t just sift through existing chemical databases. Instead, it constructs entirely new molecules using predefined building blocks and known chemical reactions. This is a subtle but important shift. Traditional generative models in chemistry often suggest structures that look promising on a screen but are practically impossible to synthesize in a lab. By design, this system avoids that trap.
In other words, it’s not just imagining molecules — it’s designing ones that chemists can realistically create.
Why MRSA Still Forces Scientists to Rethink Antibiotics
Methicillin-resistant Staphylococcus aureus (MRSA) remains one of the most challenging bacterial pathogens in modern medicine. Its resistance to multiple antibiotics makes infections difficult to treat, especially in hospital settings or chronic wounds.
This is exactly where AI-driven approaches could make a difference. The traditional antibiotic discovery pipeline is slow, expensive, and increasingly inefficient. Many known chemical classes have already been explored, leaving fewer obvious directions for researchers.
So the question becomes: where do you look next?
SyntheMol-RL approaches this problem by exploring chemical space at a scale no human team could realistically manage. Forty-six billion molecules is not just a large dataset — it’s an entirely different order of magnitude. Within that vast space, the system searches for combinations that meet multiple constraints simultaneously, something that becomes exponentially harder with traditional methods.
From Virtual Design to Real Testing: The Synthecin Case
Of course, computational predictions only matter if they translate into real-world results. This is where the study becomes particularly compelling.
After generating candidates, researchers selected 79 molecules that were not part of the training data and synthesized them in the lab. Out of these, 13 showed strong antibacterial activity against Staphylococcus aureus. Even more interesting, 7 of them were structurally distinct from known antibiotics — suggesting genuine novelty rather than incremental improvement.
Among these, one compound stood out: synthecin.
The team formulated synthecin as a topical treatment and tested it in a mouse model with MRSA-infected wounds. The results were clear — treated animals showed significantly lower bacterial levels compared to the control group.
It’s a promising signal, though it’s important to keep expectations grounded. This is still an early-stage, preclinical result. Many compounds that perform well in animal models never make it to human use.
What This Means for the Future of Drug Discovery
It would be easy to frame this as “AI has created a new antibiotic,” but that would oversimplify the reality. Synthecin is not yet a clinically approved drug, and there’s a long road ahead — including studies on mechanism of action, toxicity, safety, and scalability.
Still, the broader implication is hard to ignore.
AI in drug discovery is moving beyond simply ranking known molecules. It’s beginning to design new ones — with built-in awareness of how they can be synthesized and used in real-world conditions. That shift could significantly accelerate early-stage research, especially in areas like antibiotic resistance, where innovation has slowed in recent decades.
The real breakthrough here isn’t just one molecule. It’s the method.
And if systems like SyntheMol-RL continue to evolve, we may soon see a future where exploring billions of chemical possibilities is not an exception — but the starting point.
Source: Molecular Systems Biology






